Fibrillation Patterns Creep and Jump in a Detailed Three-Dimensional Model
of the Human Atria.
links
doi:10.1007/978-3-030-21949-9_15
abstract
Activation mapping in animal models of atrial fibrillation (AF) has shown that activation patterns can repeat for several cycles and then be followed by different changing or repetitive patterns. Subsequent clinical studies have suggested a similar type of activity in human AF. Our purpose was to investigate whether a computer model of human AF can reproduce this behavior.
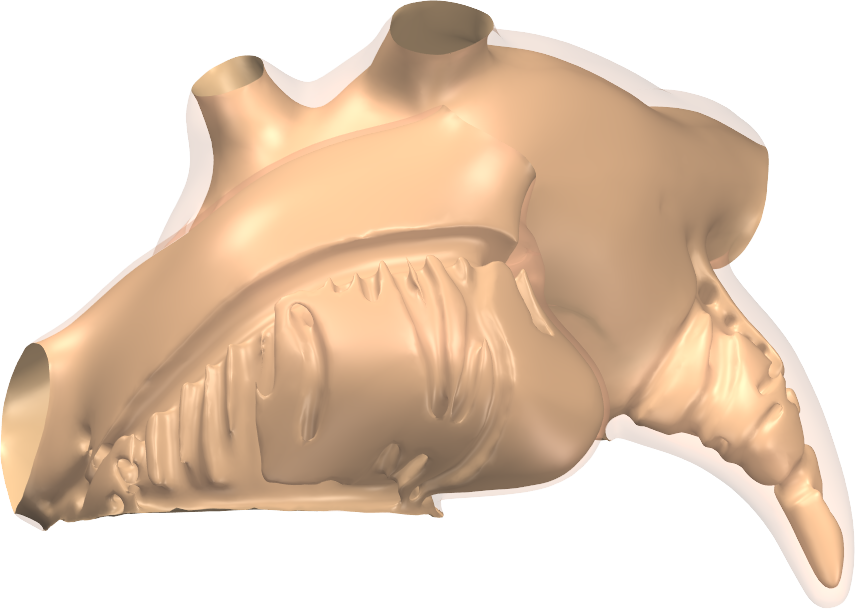
We used a three-dimensional model of the human atria consisting of 0.2-mm volumetric elements with a detailed representation of bundle structures and fiber orientations. Propagating activation was simulated over 9 seconds with a monodomain reaction-diffusion model using human atrial membrane dynamics. AF was induced by rapid pacing. Pattern recurrence was quantified using a similarity measure based on transmembrane voltage at 1000 uniformly distributed points in the model.
Recurrence plots demonstrated a continuous evolution of patterns, but the speed of evolution varied considerably. Groups of upto 15 similar cycles could be recognized, separated by rapid changes. In a few cases truly periodic patterns developed, which were related to macroscopic anatomical reentry. The drivers of non-periodic patterns could not be clearly identified. For example, a spiral wave could appear or disappear without an obvious impact on the similarity between cycles.
We conclude that our model can indeed reproduce strong variations in similarity between subsequent cycles, but true periodicity only in case of anatomical reentry.
acknowledgements
We thank Sarah Peris for the implementation of the singularity detection algorithm. This work was supported by the French National Research Agency, grant reference ANR-10-IAHU04-LIRYC. This work was granted access to HPC resources of CINES under GENCI allocation 2019-A0050307379.
the conference
This paper was presented at the 10th international meeting Functional Imaging and Modeling of the Heart, which was held at the Victoire Campus of the University of Bordeaux in June 2019. The meeting was organized by my colleagues Yves Coudière, Valéry Ozenne, Edward Vigmond, and Nejib Zemzemi, and was attended by about 120 scientists. We enjoyed keynote lectures by Hubert Cochet, Elisa Konofagou, Irene Vignon-Clementel, and Richard Gray, as well as some 30 shorter lectures on modeling of cardiac electrophysiology and mechanics, image segmentation, and inverse modeling. Sign of the time, artificial intelligence and especially deep learning was a recurrent theme. Fourty-six presented papers were published in the conference proceedings.